
This program is free software you can redistribute it and/or modify it under the terms of either: the GNU General Public License as published by the Free Software Foundation or the Artistic License. 179 Directed graphs, 10 Distribution analysis, 1011, 114, 173, 179f DNA, networks of, 91 Drawing and serialization of graphs, 153 cytoscape. perldoc Graph::Writer::CytoscapeĪll credits go to Neil Bowers for developing Graph::Writer class. You can find documentation for this module with the perldoc command. The graph, nodes, and edges can all have attributes specified, where an attribute is a (name,value) pair, with the value being scalar. Such an example is the force-directed algorithm, which draws the graph in an aesthetically-pleasing way. addgraphfromnetworkx (G2) Fully directed graphs addgraphfromnetworkx takes an argument directed that if True will ensure all edges given the directed class, which will style them with an arrow.

A lot of Apps are available for various kinds of problem domains, including bioinformatics, social network analysis, and semantic web. So, it allows you to use Graphviz file in Cytoscape. The Cytoscape JS library expects the data to be stored in a JSON file with a predefined. addedge ('separate node 1', 'separate node 2') undirected. Cytoscape is an open source software platform for visualizing complex networks and integrating these with any type of attribute data. The Cytoscape format is designed to support Cytoscape program, It simply generates the necessary files for Cytoscape. Default is None which results in the default mapping. A dictionary containing the keys ‘name’ and ‘ident’ which are mapped to the ‘name’ and ‘id’ node elements in cyjs format. A dictionary of data conforming to cytoscape JSON format. The graph must be an instance of the Graph class, which is actually a set of classes developed by Jarkko Hietaniemi. Create a NetworkX graph from a dictionary in cytoscape JSON format.
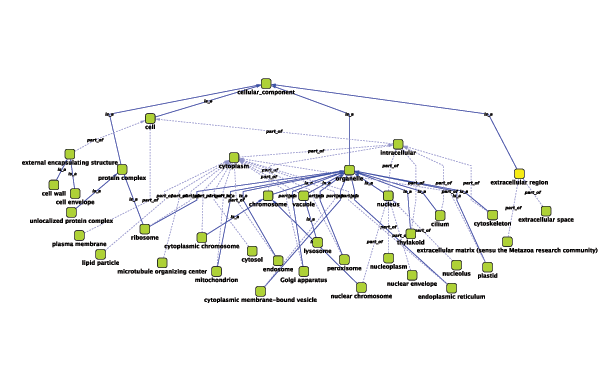
Graph::Writer::Cytoscape is a class for writing out a directed graph as Cytoscape competible input file. I will be notified, and then you'll automatically be notified of progress on your bug as I make changes. Supports rendering images of graphs on Node. Please report any bugs or feature requests to bug-graph-writer-cytoscape at rt., or through the web interface at. $writer->write_graph($graph, 'mygraph') SUBROUTINES/METHODS _write_graph AUTHOR $writer = Graph::Writer::Cytoscape->new() Graph::Writer::Cytoscape - Write a directed graph out as Cytoscape competible input file VERSION
